Scientists developed genetic tools for research of marine protists. The international 100-member team was led by Czech parasitologists.
The overlooked marine single-celled protists, which have a major impact on life on the planet, has now received greater scientific attention. One hundred and thirteen scientists from 53 laboratories from 14 countries around the world have united forces to develop genetic methods to investigate these microscopic organisms, taking ocean research a step further.
This extensive study was coordinated by molecular biologists Drahomíra Faktorová and Julius Lukeš from the Institute of Parasitology, Biology Centre (BC), Czech Academy of Sciences (CAS), and it was published on 6th April, 2020 in Nature Methods. The study was funded primarily by the Gordon and Betty Moore Foundation, EMS Program of the Marine Microbiology Initiative.
The study aimed at different types of unicellular eukaryotes. Though these microscopic sea inhabitants have been often overlooked, they play a key role of the world's ocean, moreover the life on Earth as we know it, would not exist without them. “Marine plankton produces half of the oxygen on Earth, it removes vast amounts of carbon dioxide and greenhouse gases from the atmosphere, it is crucial in ocean metabolism, as well as it is an important component of the food chain. It is surprising how little we have known about microscopic plankton so far,” says Julius Lukeš, director of the Institute of Parasitology BC CAS, whose laboratories are working on research of marine biodiversity.
The parasitologists from České Budějovice, Czech Republic, focused on the group of marine protists called diplonemids. “When I first started working with Diplonema papillatum in 2004, it was a marginal, obscure scientific topic. Only three other species of this genus were described. Today, we know that diplonemids are one of the most diverse and most abundant groups of the marine micro-world with tens of thousands of species, ” says Drahomíra Faktorová. Aiming to investigate in detail how these protists live, what they do, or how they affect the entire ocean ecosystem, scientists have focused on studying their genes. After nearly five years of research work performing hundreds of experiments, Diplonema has been successfully made a laboratory model for marine biology.

Single-celled protists Diplonema papillatum. SEM image.
“We described a precise laboratory methodology on how to perform genetic analysis on our model. Thanks to this, anyone can study the individual genes and their functions,” explains Drahomíra Faktorová.
Another 52 laboratories from 14 countries from prestigious research institutes and universities, including the famous universities of Cambridge, Harvard, Sorbonne, MIT or the University of California at Berkeley, started the same task of creating laboratory models from the other marine groups of protists. The common goal was to develop methodological protocols which will provide a path explaining how to create new protist model systems. “Not only successful methodological achievements are included in the paper. The unsuccessful approaches, which cost a lot of effort and resources, are shared too to allow others to build on our attempts. As a result, we were able to create 22 model systems out of 39 in total, while transformation of another 17 species was not achieved yet, ” summarizes Drahomíra Faktorová, who was a coordinator of the joint study.
The study represents an essential step in understanding the functioning of marine protists, their diversity, as well as family and food relationships. It will certainly contribute to the future development of the ocean research.
The study was funded primarily by the Gordon and Betty Moore Foundation, EMS Program of the Marine Microbiology Initiative.
From the sea to the bench: Marine microeukaryote model systems
Most molecular and cell biology research is performed on just a few model organisms. The most commonly studied organisms are humans, mice, yeast, worms, the zebrafish, the plant Arabidopsis and the fruit fly. Focusing on this handful of models has been meaningful and productive strategy for understanding organisms in terrestrial and freshwater environments – and during a time when obtaining any DNA sequence data was difficult and expensive. However, there is an enormous diversity of eukaryotic species in the oceans – and these are dominated by single-celled protists. Indeed, most eukaryotic diversity - both terrestrial and marine - is hidden in protists, which do not feature high on the list of model eukaryotes.
Due to their impact on human health, some parasitic protists, such as Trypanosoma brucei and Plasmodium falciparum, the causative agents of sleeping sickness and malaria, are intensely studied. However, considerably less attention has been given to free-living protists. These organisms are present in every ecosystem and have major impact on the habitability of our planet. Even within the protists, the least research has been carried out on marine species, mostly due to the lack of experimental methods – an issue exacerbated by the greater difficulty of cultivating environmentally relevant species. Yet, these are some of the most abundant organisms on planet Earth, and are fundamental to the functioning of oceanic ecosystems. Moreover, they can also be a source of major discoveries based on their unique biology.
A paper published this month in Nature Methods aims to change the way we view marine protists – and how we can study them. A consortium of 53 laboratories aimed to develop genetic tools in a range of marine protists representing the most significant groups across the eukaryotic tree of life – their collaborative efforts on this topic were supported by the Gordon and Betty Moore Foundation EMS Program of the Marine Microbiology Initiative. As a result, we can now genetically interrogate 13 species for the first time, while advanced genetic tools were developed for a further eight species and we also review an already published EMS transformation protocol in 1 species. Moreover, the paper also contains information about challenges to genetically modify 17 protist species. These unsuccessful approaches are shared to allow others to build on our attempts and to understand which methods did not work, with the hope to encourage alternative DNA delivery methods in the near future.
The collaborative effort has been summarized in a ‘Transformation Roadmap’, which provides a path explaining how to create new protist model systems. The first step is usually optimization of cultivation conditions, followed by the identification of suitable selectable markers. As long as at least one method from a wide range of approaches to deliver the exogenous DNA into the cells worked, the chances to establish a new model became high. In the next steps, various strategies were used to integrate the introduced DNA into the host genome. This introduced DNA is transcribed into RNA, and protein products are made. While there is no universally applicable protocol, and a range of conditions have been used, electroporation of linearized plasmids had the highest success rate.
These 22 genetically manipulated marine protist species now provide a springboard for functional studies of the myriad of unique and unstudied genes they carry in their genomes. With more data an unexpected pattern is emerging – namely that numerous processes and structures considered universally present and highly conserved across every eukaryotic cell is instead surprisingly divergent in these protists – challenging our textbook view about the evolution and diversity of eukaryotic life. Collectively the outcomes of this research enable scientists to delve into the secrets of the omnipresent planktonic protists, many of which can be considered the ‘plants of the ocean’ and underpin oceanic food webs and life on Earth.
NATURE METHODS WEB: https://www.nature.com/articles/s41592-020-0796-x
SCIENCE WEB: https://www.sciencemag.org/news/2020/04/new-genetic-tools-promise-unlock-secrets-microscopic-marine-life
SPRINGER NATURE BLOG: https://protocolsmethods.springernature.com/users/384116-drahomira-faktorova/posts/65306-from-the-sea-to-the-bench-marine-microeukaryote-model-systems
MOORE FOUNDATION WEB: https://www.moore.org/article-detail?newsUrlName=accelerating-the-development-of-model-systems-for-marine-microbial-ecology-through-collaboration
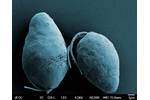
Diplonema papillatum, snímek ze...

Diplonema papillatum, snímek ze...

Eukaryotický strom života

Mikrosvět moří má zásadní vliv...














